Machine learning for carotid stenosis identification from infrared fundus images
- Joachim Behar
- 4 days ago
- 4 min read
Raphael Judkiewicz, Adi Shaked, Nomi Friedmann, Doron Hanuka, Joel Hanhart and Joachim A Behar URL: https://iopscience.iop.org/article/10.1088/3049-477X/ae5aa5
Can the eye help detect stroke risk?
A new AIMLab study explores carotid stenosis detection from routine retinal imaging
At AIMLab at the Technion, we are interested in how artificial intelligence can turn routine medical data into meaningful clinical insight. In our latest study, we explored a simple but powerful question: can images of the back of the eye help identify patients with carotid artery stenosis, a major risk factor for ischemic stroke?
Carotid artery stenosis (CAS) is a narrowing of the carotid arteries, the main blood vessels that supply the brain. It is clinically important because it increases the risk of stroke and also reflects broader systemic vascular disease. Yet screening for CAS at the population level is not routinely performed, partly because standard vascular imaging is resource-intensive and not practical for widespread use.
Our new work asks whether near-infrared retinal images, already collected during many routine ophthalmology visits, could provide a non-invasive window into this hidden vascular risk.
Why look at the retina?
The retina is one of the few places in the body where blood vessels can be observed directly and non-invasively. Because the retinal circulation is closely linked to the carotid and cerebral circulation, vascular disease upstream may leave measurable signatures in retinal structure. Prior studies have suggested that retinal vascular changes can reflect cardiovascular and cerebrovascular disease, but the link with carotid stenosis has remained underexplored and methodologically inconsistent.
Near-infrared (NIR) imaging is especially interesting in this context. These images are routinely acquired during spectral-domain OCT exams and are already part of everyday ophthalmic workflows worldwide. That means they may offer an opportunity for opportunistic screening without any added imaging burden for patients.

What we did
In this retrospective study, we analyzed 1,483 optic disc-centered NIR fundus images from 469 patients. The cohort included 121 patients with carotid artery stenosis and 348 controls. The data came from routine clinical care at Shaare Zedek Medical Center and were split into training and testing sets at the patient level to avoid leakage between sets.
We evaluated two complementary machine learning approaches:
1. A classical biomarker-based modelWe used an automated vessel segmentation pipeline to extract 28 retinal vascular biomarkers from each image, including measurements related to vessel area, tortuosity, caliber, and fractal structure. These biomarkers were then used in a logistic regression classifier.
2. A deep learning model trained on raw imagesWe also fine-tuned a self-supervised vision transformer, DINOv3-Large, directly on the NIR images, allowing the model to learn predictive patterns without explicit handcrafted features.
This dual strategy let us compare an interpretable, feature-based pipeline with a modern end-to-end deep learning approach.
What we found
The key result is encouraging: both approaches achieved an AUROC of about 0.70 on the held-out test set. Specifically, the deep learning model achieved an AUC of 0.700, and the biomarker-based logistic regression model achieved 0.702.
The deep learning model showed higher sensitivity, while the biomarker-based model showed slightly better specificity:
Deep model: sensitivity 0.73, specificity 0.60
Biomarker logistic regression: sensitivity 0.68, specificity 0.67
These numbers are not yet sufficient for standalone clinical deployment, but they do show that retinal NIR images contain detectable information associated with carotid stenosis.

Figure: ROC curve showing the performance of the Deep Learning model and the Logistic Regression with 95% CI using bootstrapping.
What makes this interesting
One of the most important findings in the paper is not only that the models worked, but where they appeared to focus. Using Grad-CAM explainability maps, we found that the deep learning model primarily attended to the retinal vasculature, rather than irrelevant image artifacts or background patterns. This supports the biological plausibility of the approach.
We also found that several vascular biomarkers were especially informative, including features related to vessel density and tortuosity. Together, these results suggest that carotid stenosis may be associated with subtle but measurable changes in retinal vascular organization.
Another important insight is that the deep learning model did not dramatically outperform the biomarker-based model. That tells us something valuable: at least in this dataset, strong vascular representations may matter more than model complexity alone.
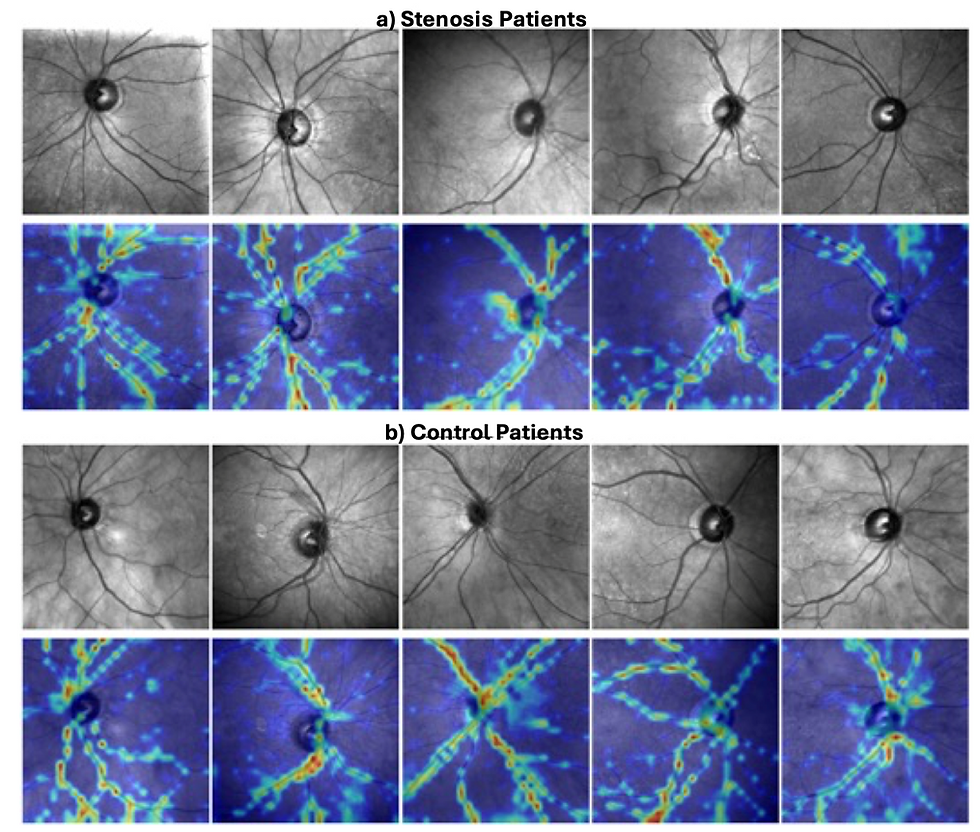
Figure: Explainability maps for Stenosis patients (a) and control patients (b). The explainability maps illustrate that the model mainly focuses on the vasculature.
Why this matters
This study is a proof of concept for a broader vision of AI-enabled medicine: using data that are already being collected in routine care to uncover risk signals that would otherwise go unnoticed. If validated in larger and more diverse cohorts, retinal NIR analysis could one day help flag patients who may benefit from confirmatory carotid imaging and more comprehensive vascular assessment.
That possibility is especially compelling because OCT and retinal imaging are already performed at massive scale. Instead of requiring new infrastructure, future tools could potentially be layered onto existing clinical workflows.
Important limitations
At the same time, this is an early-stage study, and several limitations are important.
First, the control group did not undergo mandatory carotid imaging, so some patients with undiagnosed asymptomatic stenosis may have been included as controls. Second, the dataset did not consistently include stenosis laterality or severity, which limits our ability to study dose-response effects or eye-side-specific relationships. Third, this was a single-center retrospective study, so external validation is still needed before any clinical translation can be considered.
In short: the findings are promising, but they are not a clinical screening tool yet.
Looking ahead
Our study helps define the next steps for this line of research: larger multi-center datasets, prospective validation, better carotid ground-truth labeling, and integration with clinical risk factors. In the longer term, we believe retinal AI could evolve beyond single-task prediction toward holistic ophthalmic and systemic health assessment, using routine eye exams to detect both ocular and cardiovascular disease.
At AIMLab, this is exactly the kind of challenge we care about: combining machine learning, physiology, and real-world clinical data to build tools that are both scientifically rigorous and clinically meaningful.
Publication
Judkiewicz, R., Shaked, A., Friedmann, N., Hanuka, D., Hanhart, J., & Behar, J. A.Machine learning for carotid stenosis identification from infrared fundus images.Accepted manuscript, Machine Learning: Health. https://iopscience.iop.org/article/10.1088/3049-477X/ae5aa5




Comments